 |
Welcome to the laboratory of Ruohe Yin |
 |
Welcome to the laboratory of Ruohe Yin |
In our laboratory we investigate how plants sense and respond to light signals in a way to improve their adaptation to the dynamic environment. Currently we focus on the function of UV-B photoreceptor UVR8 (Rizzini et al., 2011). Several key functions of UVR8 have been revealed including regulation of seedling photomorphogenesis, UV-B acclimation and UV-B stress tolerance, secondary metabolism, phototropism, root development and more. UV-B induces UVR8 homodimers dissociation into monomers to trigger UV-B signaling. UV-B induces UVR8 translocation into nucleus (Kaserli et al., 2007). In the nucleus, UV-B-activated UVR8 monomers regulate the expression of a group of genes via different mechanisms including inactivation of E3 ligase COP1 to release its degradation targets and direct association with several transcription factors and more (Jenkins 2017; Liang et al., 2019; Podolec et al., 2021). It
Previously, we reported that UV-B-induced UVR8 accumulation in nucleus requires COP1 with unknown mechanisms (Yin et al., 2016) (Fig. 1). Based on the observations that UV-B induces UVR8-COP1 interaction in plant cell and COP1 contains a bipartite NLS, we use a combination of molecular genetics, cell biology and biochemistry to investigate the molecular mechanisms by which COP1 regulates UVR8 nuclear accumulation. We also screen for novel protein candidates that participate in the regulation of UV-B-induced UVR8 nuclear-cytoplasmic distribution.
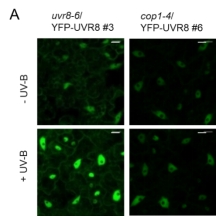
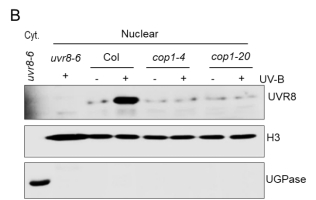
Fig. 1 COP1 is required for UV-B-induced UVR8 protein accumulation in nucleus.
The bZIP transcription factor ELONGATED HYPOCOTYL5 (HY5) plays key roles in diverse light signaling pathways in plants and acts downstream of multiple photoreceptors. A subgroup of UVR8-regulated UV-B target genes are regulated by HY5 and its homolog HYH (Brown et al., 2005; Brown & Jenkins, 2008; Favory et al., 2009). Tomato genome contains a single HY5 (SlHY5). Transcript of SlHY5 is induced and then rapidly attenuated by UV-B, suggesting the existence of unknown negative regulation of SlHY5 transcription in prolonged UV-B treatment (Fig. 2), which is critical to halt UV-B signaling. HY5 was shown to be required but not sufficient to activate its gene transcription (Abbas et al., 2014; Binkert et al., 2014). Therefore, neither the transcription factors that activate HY5 transcription nor the mechanism for its attenuation during UV-B signaling is known. We aim to identify the factors that activate HY5 transcription and detailed mechanisms for the attenuation of HY5 transcription in the prolonged UV-B treatment. We found that SlHY5 transcript levels were induced by UV-B, peaking after 1 h and then declined rapidly in wild-type tomato seedlings (Fig. 2).
RUP1 and RUP2 are key negative regulators in the UVR8 pathway (Grube et al., 2010). They interact with UVR8 and inactivate UVR8 by promoting the reassociation of UVR8 monomers (active) to form inactive UVR8 homodimers (Heijde et al., 2013). In tomato a single RUP protein is present with similar functions in UV-B signaling (Zhang et al., 2021). Interestingly, UV-B-induced SlHY5 transcription was higher in the slrup mutant than in WT (Fig. 2). Notably, the attenuation of UV-B-induced SlHY5 transcription was delayed by 1-3 h in the slrup mutant relative to WT (Fig. 2). Thus, SlRUP participates in, but is not sufficient for the negative regulation of SlHY5 transcription in response to UV-B.
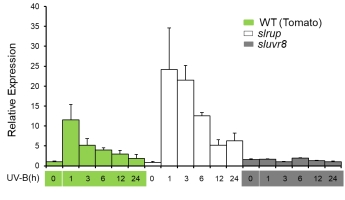
Fig. 2 Transcript of SlHY5 is induced by UV-B transiently in tomato.
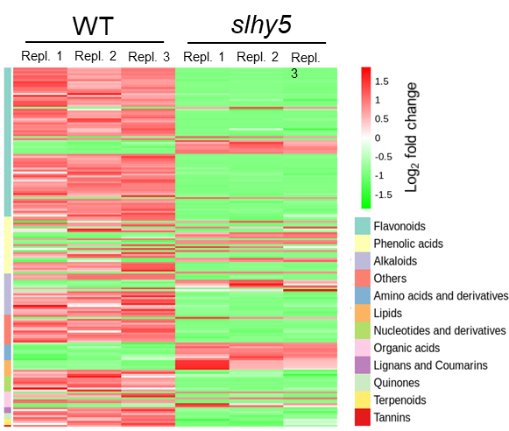
In agriculture, low light can induce exaggerated stem elongation and reduce yield and fruit quality. We take tomato as a model to investigate how crop plants respond to light. We found that tomato fruit metabolism is greatly altered in slhy5 mutant (Fig. 3). Our long-term goal is to improve crop tolerance to low light for better yield and fruit quality. we also investigate tomato and Arabidopsis species-specific UV-B signaling mechanisms.
Fig. 3 Fruit metabolism is strongly altered in slhy5 mutant.